Method reveals how proteins read RNA messages
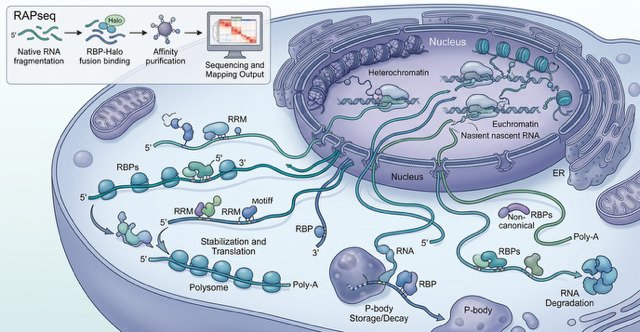
Researchers at Karolinska Institutet have developed a technique that shows how proteins interact with RNA, the molecule that carries genetic instructions inside our cells. The study, published in Nucleic Acids Research, offers new insight into an important layer of gene regulation that can lead to better understanding of disease mechanisms.
RNA-binding proteins (RBPs) decide when RNA messages are used, how long they last, and how much protein is produced from them. However, mapping these interactions has long been technically challenging. In the new study, the research team presents RAPseq, a method that can systematically chart how RBPs bind to RNA across the whole transcriptome.
Using RAPseq, the researchers showed that both well‑studied and lesser‑known RBPs bind to RNA in specific patterns. They also revealed how cancer‑related mutations change these interactions, and identified differences in RNA recognition across vertebrate species.

“RAPseq allows us to capture how RNA‑binding proteins manage genetic messages, which helps us understand what happens when these regulatory processes are altered in disease”, says Principal Researcher Claudia Kutter at the Department of Microbiology, Tumor and Cell Biology at Karolinska Institutet.
Contribute to a better understanding of disease mechanisms
The method can be used to study previously uncharacterized proteins, investigate disease‑associated variants, and identify disrupted RNA networks in conditions such as cancer. It may also support the development of future strategies to influence harmful RNA–protein interactions.
“By making it easier to study these regulatory events, we hope the results will contribute to better understanding of disease mechanisms and improved approaches to diagnosis and treatment,” says Claudia Kutter.
The team now aims to apply RAPseq to additional mutations and integrate the method with functional cellular experiments to identify the most biologically relevant RNA‑protein interactions.
The work was led by Principal Researcher Claudia Kutter at the Department of Microbiology, Tumor and Cell Biology, together with first authors Riccardo Mosca, Carlos Gallardo‑Dodd and Qun Li.
The study was supported by the Knut & Alice Wallenberg foundation, Cancerfonden and VR. See the study for any reported conflicts of interest.
Publication
"RAPseq enables large-scale identification of RBP-RNA interactions and reveals essentials of post-transcriptional gene regulation", Mosca R, Gallardo-Dodd CJ, Li Q, Sommerauer C, Šidiškis J, Søndergaard JN, Kutter C. Nucleic Acids Research, online 27 February 2026, doi: 10.1093/nar/gkag090.
