Synthetic antibody can block SARS-CoV-2 infection
By screening hundreds of synthetic antibodies, researchers at Karolinska Institutet in Sweden and EMBL Hamburg in Germany have identified an antibody that may prevent the new coronavirus from infecting human cells. The study, which is published in the journal Nature Communications, also shows how antibodies can be quickly produced in the event of future pandemics.
On the surface of SARS-CoV-2 are spike proteins that give coronaviruses their characteristic appearance and help them to infect cells. These proteins use their three finger-like projections, called receptor-binding domains, to bind to the surface protein ACE2 on human cells. Once SARS-CoV-2 has bonded with ACE2, the membrane of the virus melds with the host cell membrane, allowing the viral material to enter and infect the cell.
Antibodies that prevent the spike proteins from binding to the host cell can thus block SARS-CoV-2 infection. Fragments of antibodies, called nanobodies, that occur naturally in camelidae, have previously been produced to prevent SARS-CoV-2 from entering cells, but the development process is relatively time consuming.
Cheap to produce

“In a collaboration between EMBL Hamburg and Karolinska Institutet, we’ve managed to develop nanobodies, so-called sybodies, which are not only tiny but also extremely stable and relatively simple and cheap to produce,” says Martin Hällberg, researcher at the Department of Cell and Molecular Biology at Karolinska Institutet.
Dr Hällberg’s research group has been working closely with Ben Murrell’s and Gerald McInerney’s groups at KI and Christian Löw’s group at EMBL Hamburg in Germany to find sybodies able to block SARS-CoV-2 infection using a newly developed technical platform. They found that some of the hundreds of synthetic antibodies produced, especially one called Sybody 23, proved extremely effective in blocking the binding process between the viral spike proteins and the human surface protein ACE2.
Determining the structure
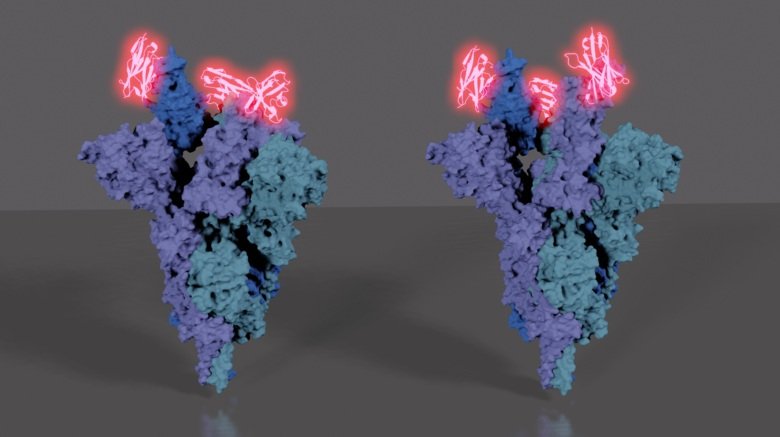
The researchers then employed electron cryomicroscopy (cryo-EM) at KI’s recently opened 3D-EM facility to gain an understanding of how the synthetic nanobody blocks SARS-CoV-2 by determining the structure of the spike proteins bound to Sybody 23.
The spike protein’s finger-like projections can be either “up” when binding to ACE2 or “down” when hiding from the human immune system. The structural studies showed that Sybody 23 binds to both these states, effectively blocking the areas where the binding process would normally take place on ACE2. The studies also showed that spike proteins bound to Sybody 23 adopt two different conformations.
Can be useful in a future pandemic
This structural insight will make it easier for the researchers to design combinations of sybodies able to bind to different areas of the SARS-CoV-2 virus’s spike proteins. They hope then to be able to improve the effectiveness of the antibodies and make it harder for the virus to avoid neutralisation by mutating.
“We show that the use of synthetic antibody libraries combined with structural studies presents good opportunities to quickly produce efficacious therapeutic antibodies, which can also be useful against new viruses in a future pandemic,” says Dr Hällberg.
The study was supported in part by grants from the EU (Horizon 2020) and the Swedish Research Council.
Publication
”Selection, biophysical and structural analysis of synthetic nanobodies that effectively neutralize SARS-CoV-2”. Tânia F. Custódio, Hrishikesh Das, Daniel J Sheward, Leo Hanke, Samuel Pazicky, Joanna Pieprzyk, Michèle Sorgenfrei, Martin Schroer, Andrey Gruzinov, Cy Jeffries, Melissa Graewert, Dmitri Svergun, Nikolay Dobrev, Kim Remans, Markus A. Seeger, Gerald M McInerney, Ben Murrell, B. Martin Hällberg, Christian Löw. Nature Communications, online 4 November 2020, doi: 10.1038/s41467-020-19204-y.
