New 3D imaging method for tumour samples to be used for studying COVID-19
Researchers at Karolinska Institutet in Sweden have developed a new method for analysing intact tumour samples in 3D on a molecular level. The study is a collaboration with Karolinska University Hospital and is published in Nature Biomedical Engineering. Now the researchers are planning to use the method to study tissue samples from COVID-19 patients in the hope that it will provide information on where and how the new coronavirus attacks different organs.
Cancer is caused when vital molecular mechanisms in cells stop operating normally. To better understand these pathological processes, scientists need to study what occurs in tumour cells at an RNA and protein level. RNA acts as a messenger between the genes and the proteins that they code for, and there are different kinds of RNA involved in cancer.
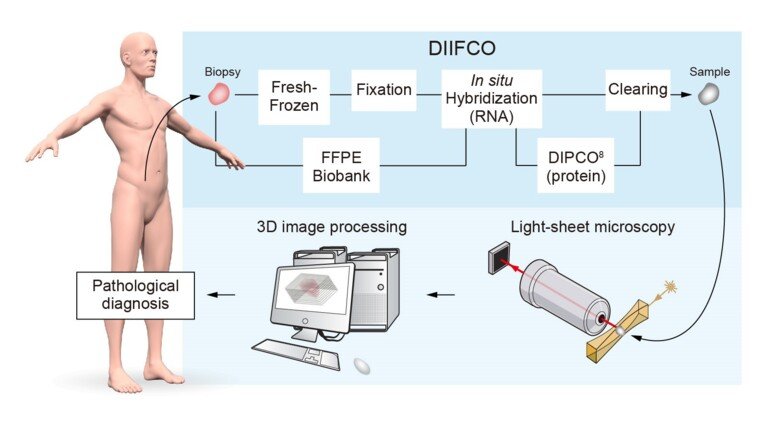
The researchers have applied a new imaging technique, which is used in basic research, to study intact – rather than sectioned – human tumour tissue. The technique involves making the tissue transparent, staining different kinds of RNA and proteins, and then imaging the tissue in three dimensions using an instrument called a light-sheet microscope. Doing this, it is possible to determine where in the tumour there are abnormal amounts of RNA or protein. The method, called DIIFCO, facilitates the study of cancer-related mechanisms and enables more exact diagnoses of tissue samples from cancer patients (see figure 1).

More exact cancer diagnoses
“For years, clinicians have used the labelling of specific proteins to diagnose tissue samples from tumours,” says principal investigator Per Uhlén, professor at the Department of Medical Biochemistry and Biophysics. “Being able to specifically stain RNA in intact tumour samples opens up entirely new opportunities for clinical specimen diagnosis.”
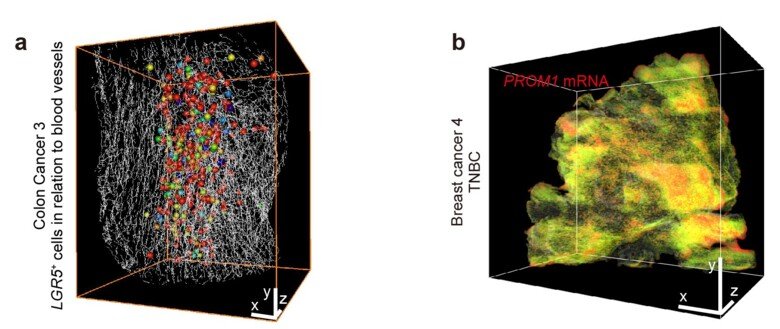
Working with pathologists and surgeons at Karolinska University Hospital, the researchers studied digital 3D images of tumours using advanced image processing software. In this way, they managed to identify each individual cell of a tumour and to analyse how the cells cluster as well as how specific cell populations arrange themselves in relation to the tumour’s blood vessel structure (see figure 2a). In another experiment, they studied the presence of so-called cancer stem cells in a highly aggressive form of breast cancer by specifically staining PROM1 RNA, a biomarker for cancer stem cells. Cancer tumours contain a very small number of cancer stem cells, which are thought to play a significant part in cancer formation. The DIIFCO method revealed small clusters of cancer stem cells deep inside the tumour samples from breast cancer patients (see figure 2b).
COVID-19 samples next
Using the DIIFCO method, scientists can study RNA and proteins at a high resolution in all human tissue. The researchers are now planning to use their method to study tissue samples from COVID-19 patients. PCR analyses for detecting the RNA from the SARS-CoV-2 virus are frequently used in hospitals to test if patients have COVID-19, and the researchers now hope that the new imaging technique will be able to detect exactly where in the tissue the viral RNA is.
“Ours is one of the few labs in the world able to study intact tissue samples from humans,” says Professor Uhlén. “This is particularly difficult when the tissue comes from elderly patients. I believe and hope that our method can provide important answers to where and how the new coronavirus attacks different organs.”
The study was financed with grants from the Swedish Cancer Society, the Swedish Childhood Cancer Foundation, the Swedish Brain Fund and the Swedish Research Council.
Publication
“Three-dimensional single-cell imaging for the analysis of RNA and protein expression in intact tumour biopsies”. Nobuyuki Tanaka, Shigeaki Kanatani, Dagmara Kaczynska, Keishiro Fukumoto, Lauri Louhivuori, Tomohiro Mizutani, Oded Kopper, Pauliina Kronqvist, Stephanie Robertson, Claes Lindh, Lorand Kis, Robin Pronk, Naoya Niwa, Kazuhiro Matsumoto, Mototsugu Oya, Ayako Miyakawa, Anna Falk, Johan Hartman, Cecilia Sahlgren, Hans Clevers, Per Uhlén. Nature Biomedical Engineering, online 29 June 2020, doi: 10.1038/s41551-020-0576-z.
