Newsletter for staff and affiliates at CMB - March 2026

Newsletter for staff and affiliates at the Department of Cell and Molecular Biology, March 2026.
A few introductory words from the chairman
Dear colleagues,
I hope 2026 has started well for all of you!
At “central” CMB level we are busy with the normal activities, and also prepare for the final steps in an ongoing evaluation of the research performed at campus Solna and Syd. The evaluation includes a meeting between CMB advisory board and an external panel of very prominent scientists, who also will present their research in the Klein lecture hall 9th of March. Don’t miss the opportunity, you are most welcome to attend. In this letter you will find information on new CMB colleagues, a report from our conference, an AI column by Olof Nordenstorm in Avlant’s team and more.
Looking forward to yet another exciting year as your head of department.
News from the Head of Administration
As the winter months slowly give way to spring, the days are once again becoming noticeably lighter. The first indications of spring—slightly longer afternoons, milder air and even the occasional glimpse of the sun after its long absence—signal that the season is beginning to shift.
Our administration team continues its work with dedication and remains available to support you with any inquiries you may have, whether large or small. Please feel free to contact us whenever assistance is needed.
We would like to take this opportunity to inform you about the upcoming KI employee survey, which is planned to be distributed by email at the end of March. In addition, we would like to remind everyone of the current guidelines for expenses and representation. You will find more information about the employee survey and the guidelines a little further down on the page. Should you have any questions, our administration team is available to assist.
We wish everyone a pleasant and inspiring season ahead!
CMB Confrence 2025
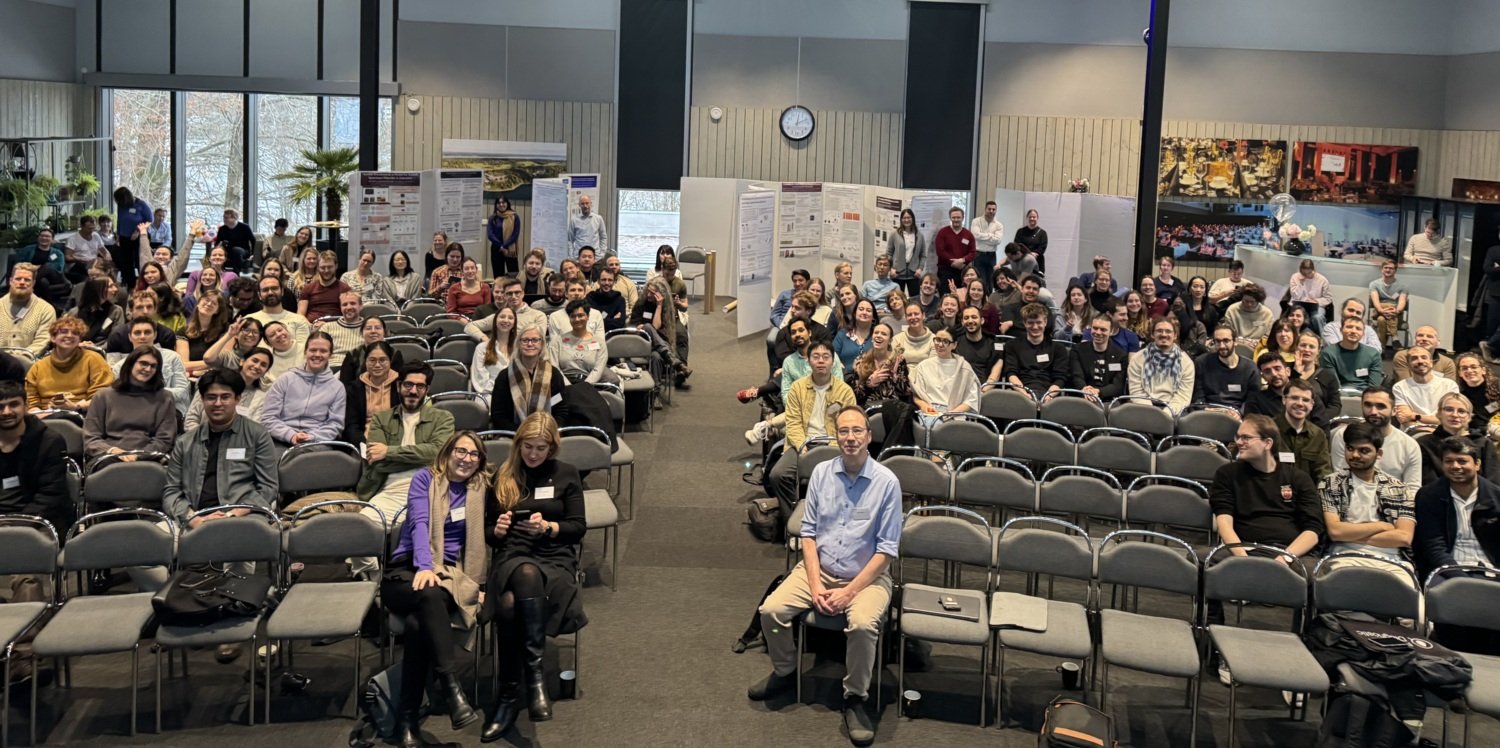


The CMB department conference held November 27–28, 2025 on Lidingö gathered students, postdocs and principal investigators for a one-and-a-half-day program on advancing cell, molecular and developmental biology.
The conference combined scientific presentations and team-building activities. Highlights of the program included research talks from all CMB groups, poster flash talks, and a panel discussion on departmental life.
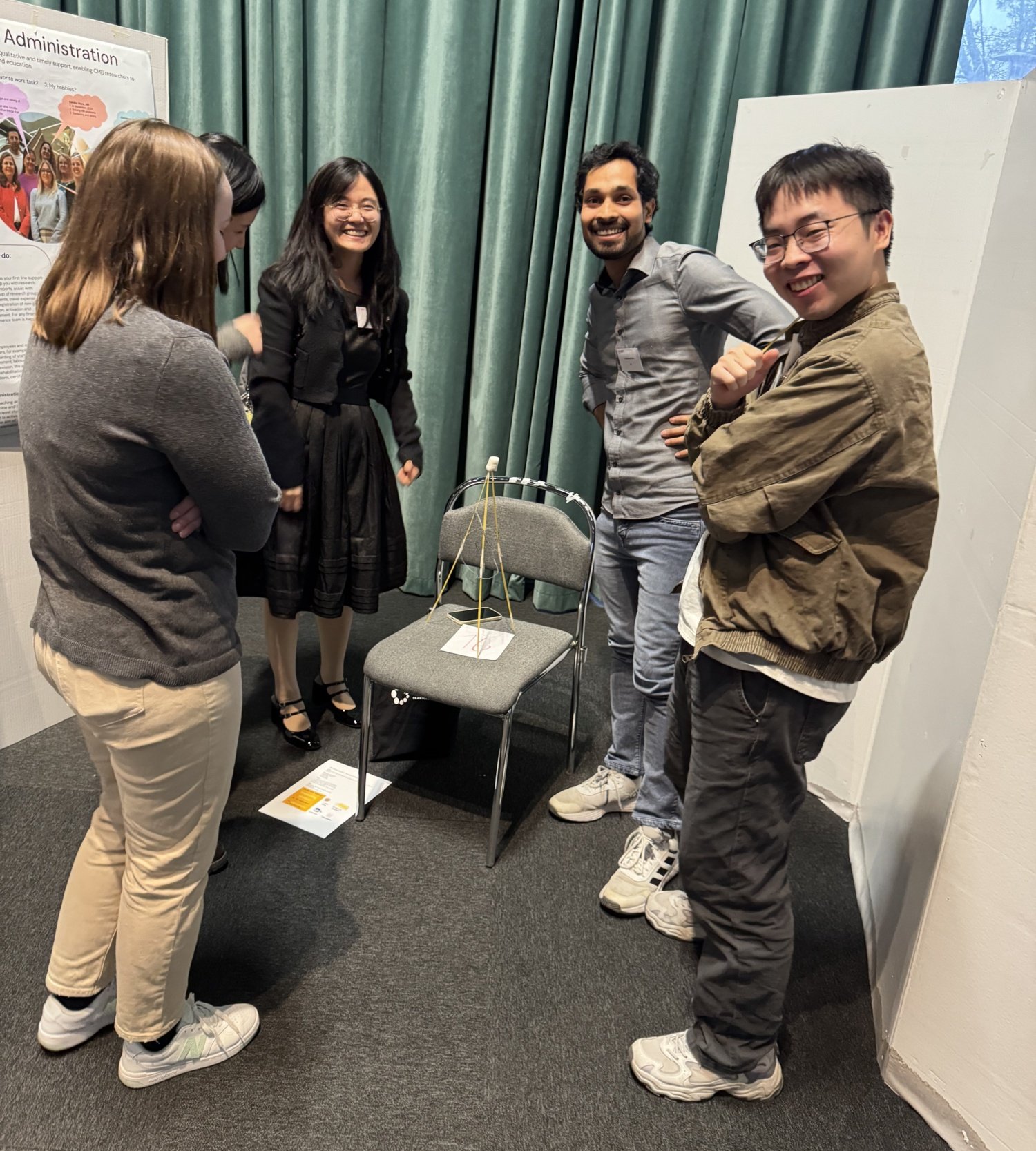
Marshmallow and spagetti
Randomized groups, more than 150 participants, and the task to build the tallest, freestanding structure with the marshmallow on top. The winning team: a team of five students and postdocs, no professors involved, and a lot of iterative “revise & resubmit".
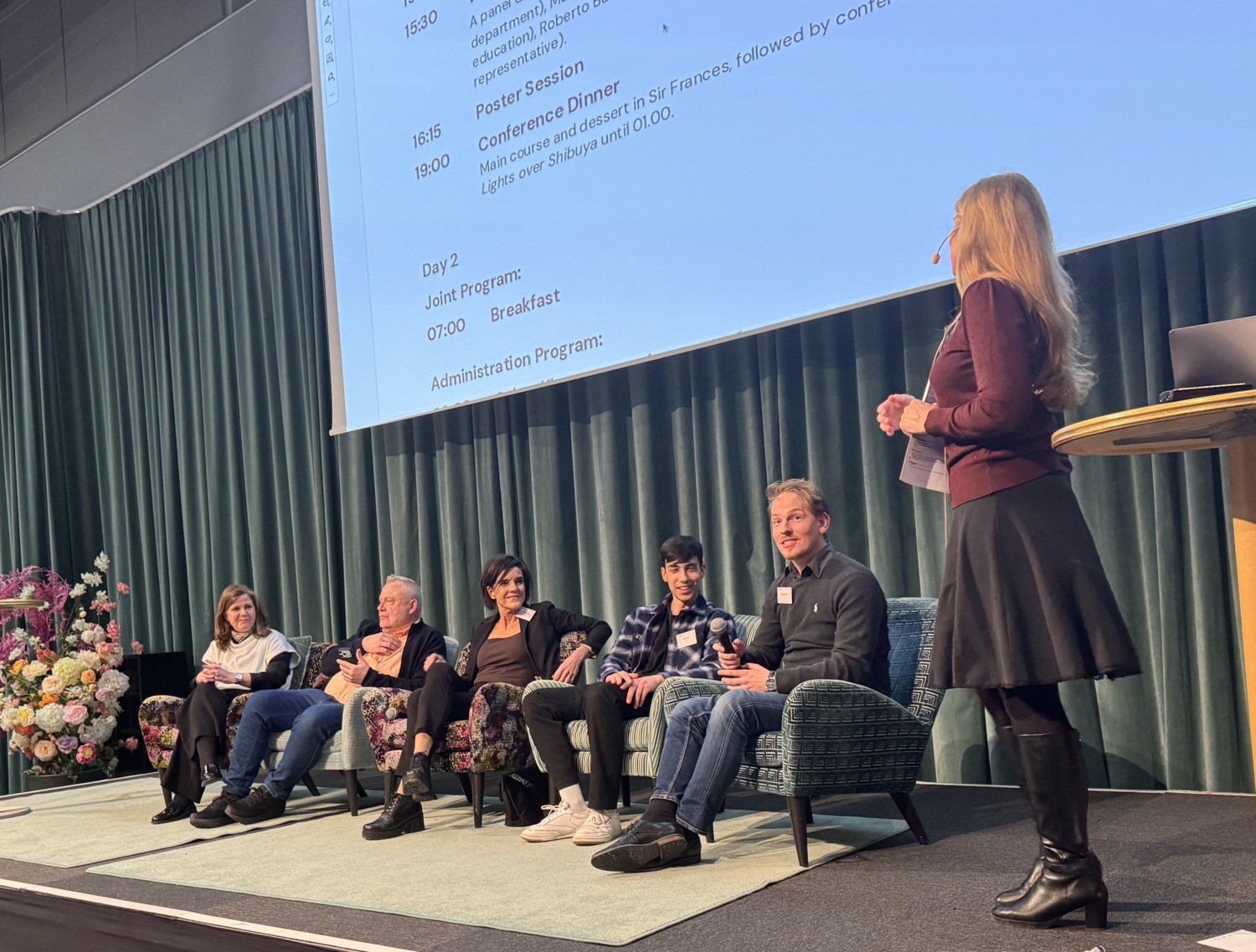

Panel discussion
With Emma Andersson (moderator), Camilla Björkegren, Margaret Ulander, Matti Nikkola, Roberto Ballarino, Ole Unseld brought together diverse perspectives on a wide range of topics including AI, community building at CMB for a lively and honest exchange. The conversation was dynamic, nuanced, and could have continued well beyond the allotted time - always a good sign.

Exchange and community building
The evening followed by a social dinner and networking activities under the theme *Northern Lights over Shibuya*.
Overall, the conference was regarded as a successful and inspiring departmental gathering that highlighted the breadth of research and strengthened connections across groups.
Post‑conference feedback emphasized appreciation for the balanced program, opportunities to discover colleagues’ work, and informal discussions that supported collaboration. Participants enjoyed the mix of speakers, interactive elements, and organization. Suggestions for future events included allocating more time for informal networking and adjusting the poster‑session timing to encourage broader participation and discussion.
The organizing team Michael Ratz, Emma Andersson, Avlant Nilsson, Roberto Ballarino, Ole Unseld, Mia Sjöblom thanks all participants for their engagement and feedback, which will guide improvements and help strengthen the collaborative spirit of the department in future events.

The winners
Winners of the marshmallow challenge group 16: Hrishikesh Das, Zhiyu Hao, Xuechun Xu, Zikang Zhang and Aleksandra Zielinska with a 68 cm high free standing spaghetti tower.
The talk prize winners were: Niels Krämer, Jeff Mold, and Valentina Furlanetto.
The poster prize winners were: Simon Perrin, Rosa Marís López Cabezas, and Evanthia Iliopoulou.
AI column
Will we see a virtual AI cell before 2030?
Just a few years ago, a comprehensive computational model of a living cell seemed far-fetched. Today, a race is underway to achieve it. Perspective papers in Cell (1), community initiatives like the Virtual Cell Challenge (2), and national and corporate investments, including Sweden’s Alpha Cell programme (3), signal a concerted push toward this goal. DeepMind, the creators of AlphaFold, have also entered the field with both a clear intent to create a virtual cell as stated in multiple interviews with Demis Hassabis (4) but also with the launch of their new model Alpha Genome (5).
Several converging forces make this moment plausible. Omics data have exploded in scale and diversity, from high-throughput perturbation screens (6) and spatial transcriptomics to multimodal single-cell profiling and high-content imaging. Deep learning has advanced rapidly alongside GPU improvements. Given this unprecedented advancement a full cell model in 2030 still seems unlikely. Cells are highly complex, heterogeneous, and context-dependent, and many data dimensions remain sparse, additonaly batch effects complicates integration of diverse data sources. Nevertheless, innovative methods continue to emerge that tries to tacle these challenges (7-9). It is clear that a race is on, and if the launches of both ChatGPT and AlphaFold have taught us anything, it is that deep learning is a technology capable of delivering unexpected breakthroughs.
- https://www.cell.com/cell/fulltext/S0092-8674(24)01332-1
- https://virtualcellchallenge.org/
- https://www.scilifelab.se/alpha-cell/
- https://lexfridman.com/demis-hassabis-2-transcript/
- https://www.nature.com/articles/s41586-025-10014-0
- https://www.biorxiv.org/content/10.1101/2025.02.20.639398v1
- https://www.nature.com/articles/s41592-025-02877-y
- https://www.nature.com/articles/s42256-025-01150-3
- https://www.nature.com/articles/s42256-025-01165-w
Lab news
Rickard Sandberg awarded Hans Wigzell Research Foundation's scientific prize of SEK 1 million
Hans Wigzell Research Foundation's scientific prize for 2025, worth SEK 1 million, has been awarded to researcher and professor Rickard Sandberg for his groundbreaking research into RNA and how RNA activation in individual cells is initiated and controlled.
A long-sought goal in medical research has been to be able to study RNA molecules in individual cells. Rickard Sandberg has developed new methods that make this possible, and in his research, Sandberg has also made several unexpected and dramatic discoveries about how RNA activation in individual cells is initiated and controlled.
“Sandberg has made a global pioneering effort in developing techniques that make it possible to study RNA molecules in individual cells. Sandberg's research has given the research community new exciting tools for analyzing RNA from individual cells from different tissues in both health and disease,” says Hans Wigzell.
Welcome new recruits
Nico Dantuma's lab

Simon le Goupil
Postdoctoral ResearcherI completed my Ph.D. in Cancer Biology at the Oncogenesis Stress Signaling Lab (France), where I focused on endoplasmic reticulum proteostasis and the protein-protein interactions of the IRE1 sensor. Eager to advance my career in academia, I joined Nico Dantuma’s research group in the spring of 2026. My current PostDoc research investigates the copper ionophore-dependent regulation of the proteasome and autophagy. I believe that better understanding copper signaling mechanisms will allow us to design novel therapeutic strategies for diseases driven by protein degradation dysregulation, such as cancer and neurodegeneration.
Avlant Nilsson's lab
Beate Gündel
Postdoctoral ResearcherBeate Gündel with a PhD from KI on in vitro modeling of tumor-stroma-crosstalk in pancreatic cancer.
She will be working with high throughput quantification of ligand responses in pancreatic cancer and stroma cells.
Michael Ratz's lab
Dheeraj Chandra Joshi
Postdoctoral ResearcherI completed my Ph.D. from CSIR-Institute of Genomics and Integrative Biology, New Delhi, India, with a focus on identifying the inherited long non-coding RNAs and understanding their role in gene expression during early zebrafish development. After my Ph.D., I joined Michael Ratz’s group on an interdisciplinary project with Johanna Lanner (FyFa, KI) that focuses on understanding the mechanisms of motor impairments in autism using zebrafish as an in vivo model.

Jatin Sharma
Research AssistantI completed my MSc Neuroscience at the University of Bremen with my thesis work focusing on iPSC-neuronal models of neurodegeneration at DPAG, Oxford. My background includes a range of tools studying cognition and biology, ranging from fMRI/fNIRS to in-vivo chemogenetic/optogenetic approaches in rodents, and further in-vitro pharmacological methods. I joined Michael Ratz's group, where I am currently exploring the utility of zebrafish as a model for autism by profiling their behavioral and molecular phenotypes, with the goal of gaining insight into the mechanisms underlying ASD.

Oumeng Yuan
Research AssistantI studied Biomedicine at the University of Toronto, Canada and at Karolinska Institute. I joined Michael Ratz’s group as a research assistant in August 2025. My current research focuses on studying early neurodevelopment by tracking barcoded lineages in vivo and integrating spatial transcriptomics to map how cellular fate and circuit organization emerge during development.

Lucia Adam
Master studentI have studied Neuroscience at the University of Edinburgh, Scotland, and I am currently completing my master’s degree in the Biomedicine program at Karolinska Institute. I joined the Ratz group in January 2026 to investigate the effects of cell-adhesion molecule gene knockdown on neuronal gene expression and circuit maturation.

Nikolitsa Kritsioni
Master studentI studied Chemistry at the University of Patras, Greece and I am currently pursuing a Master’s degree in Neurochemistry at Stockholm University. During my Master’s, I became especially interested in brain development and joined Michael Ratz’s lab for my Master thesis in September 2025. My work focuses on monosynaptic rabies tracing to map emerging connections in the mouse cortex, contributing to our understanding of mammalian brain development.
Kirsty Spalding's lab
Morteza Aslanzadeh
BioinformaticianI am a bioinformatician with a background in computational and molecular biology. I completed my PhD in quantitative RNA biology, focusing on RNA regulation, small RNA biology, and large-scale sequencing data analysis. My work involves developing reproducible analysis pipelines and interpreting bulk and single-cell transcriptomic data. In the Spalding lab, I am interested in applying computational approaches to better understand human adipose tissue biology and its role in health and disease.
New at CMB and want to participate in teaching? Talk to Matthew Kirkham, Arne Lindqvist, Lena Ström or Matti Nikkola.
Doctoral education
Welcome to newly admitted doctoral students
- Hallgerður Kolbeinsdóttir, main supervisor Laura Baranello (4 August 2025)
- Xinhe Xing, main supervisor Avlant Nilsson (4 August 2025)
- Caroline Lang, main supervisor Kirsty Spalding (5 August 2025)
- Jonas Sharma, main supervisor Rickard Sandberg (1 September 2025)
- Sam Kindl, main supervisor Maria Kasper (22 September 2025)
- Francesca Binando, main supervisor Kristian Jeppsson (1 October 2025)
- Zikang Zhang, main supervisor Rickard Sandberg (3 December 2025)
Doctoral theses defences
Congratulations to
Benjamin Dedic, who successfully defended his thesis on December 5th 2025: "The Role of the Insulin Receptor in Mature Human Adipocytes".
Hao Yuan, who successfully defended his thesis on January 22nd 2026: "Efficient integration of human skin single-cell RNA sequencing data"
Rebecca Hamilton, who successfully defended her thesis on February 20th 2026: "From Survival to Long-Term Health: Understanding and Improving Outcomes for Children Undergoing Congenital Heart Surgery".
Communication
Employee survey
From 30 March to 17 April, a new round of the employee survey will be conducted to provide an update on our workplace environment. The survey is an important part of KI’s systematic work environment efforts and a way to follow up on working environment, wellbeing, and leadership.
The survey is distributed via email by KI’s partner Agerus, who manages all responses and ensures anonymity. The survey is available in both Swedish and English and will remain open for three weeks, from 30 March to 17 April.
The survey consists of two parts. The first covers the employee Net Promoter Score (eNPS), which measures how likely you are to recommend KI as a workplace. The second part comprises four index areas: Employee engagement, leadership, organization, and goals & strategies.
For more information: Employee survey 2026 | Staff Portal
Guidelines on entertainment, gifts and certain benefits in kind.
Karolinska Institutet (KI) must, as a public authority, manage state funds responsibly. This applies particularly to representation, gifts, and staff benefits. These guidelines are directed at employees who use funds for representation, gifts, and certain staff benefits. The purpose of the guidelines is to clarify the difference between external and internal representation and when and to what extent business entertainment is appropriate.
Guidelines on entertainment, gifts and certain benefits in kind | Staff Portal
Ordering IT services at KI
If you have questions about how to order software licenses, servers and data storage, generative AT et cetera, you can read more about the various services offered by the IT Office at KI:
Order IT and telephony services | Staff Portal or contact KI Helpdesk for more specific questions: Additional IT support services | Staff Portal
New instrument
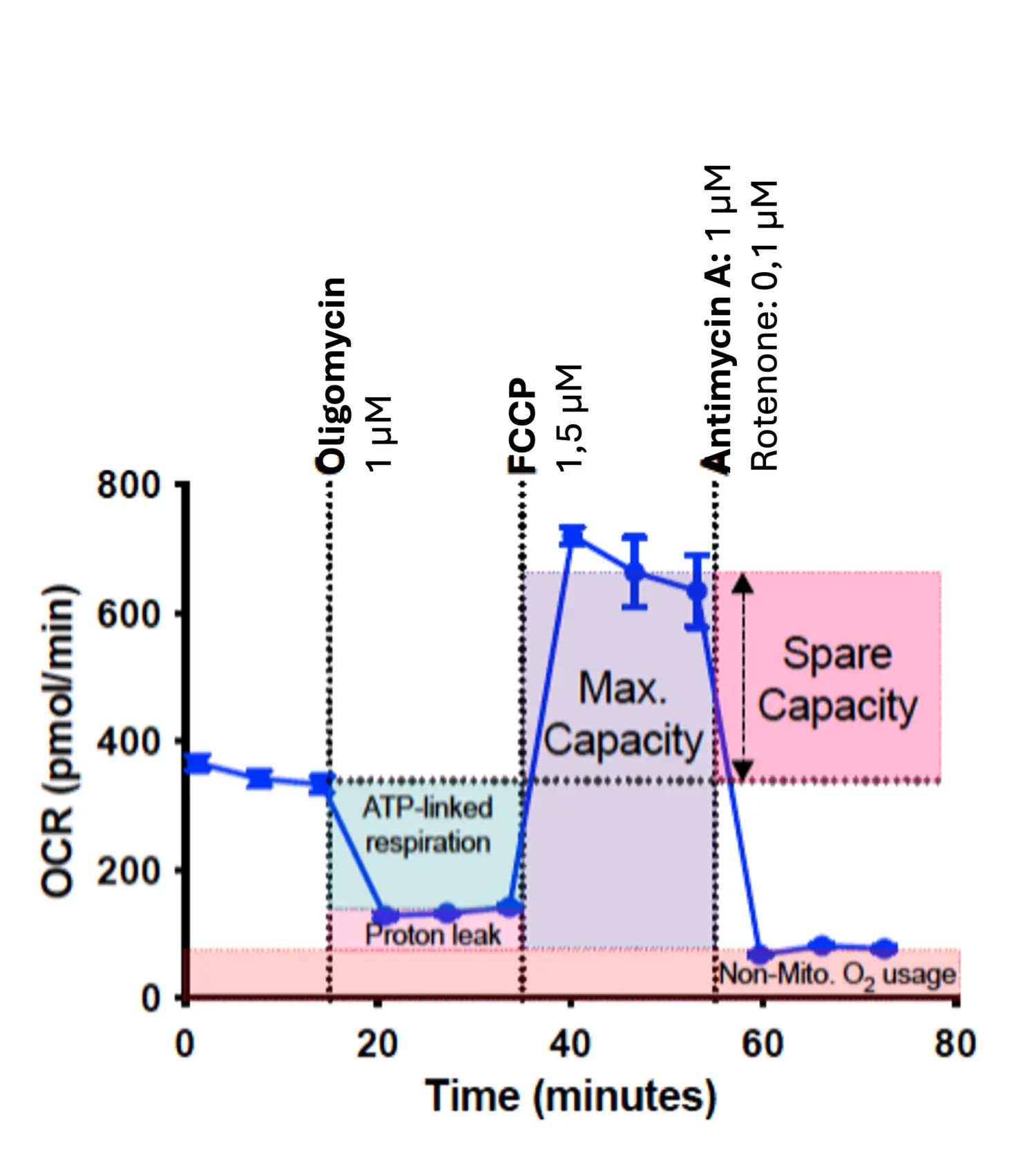
Seahorse XFe96 Analyser
This instrument is now available as part of a core facility in Biomedicum for all users who are interested in metabolic characterisation of their cells. The Seahorse can be used for a variety of applications, such as characterising disease models, identifying drug targets, validating target effects on cellular function, and determining drug safety.
Seahorse XFe96 Analyser available at Biomedicum (C0528).
For more information or for booking an introduction contact: seahorse-biomedicum@ki.se
Seminars
The Avlant Nilsson group is happy to invite you to two seminars on the theme of Biologically Informed Neural Networks, Friday March 13 at 10am in room Ragnar Granit https://news.ki.se/calendar/biologically-informed-neural-networks. The talks will highlight how integrating prior biological knowledge with modern optimization and neural network approaches enables more interpretable, predictive, and scalable models of disease and omics data. Join us for an engaging discussion!
Your profile page in KI RIMS

Make sure to review and update your personal profile page in KI-RIMS. Your profile page is usually the first hit when searching for your name on ki.se or when searching via other search engines.
