Circular RNAs in Hedgehog signaling activation and Hedgehog-mediated medulloblastoma tumors
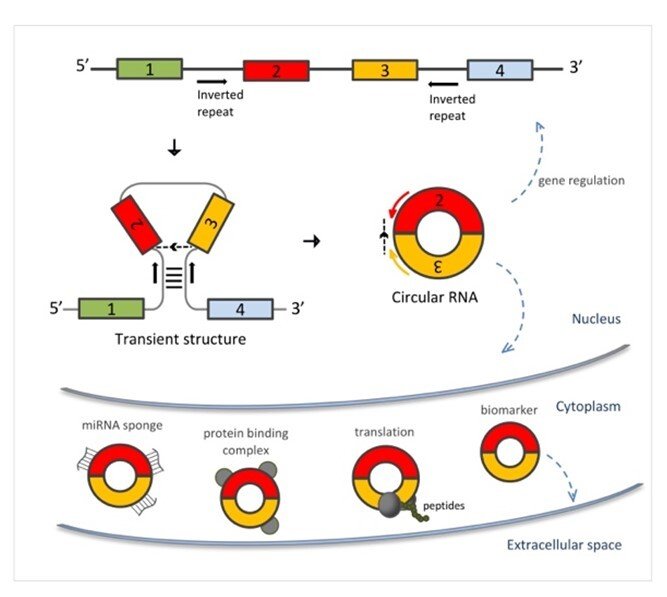
Circular RNAs’ role in cancer initiation and progression has gain a lot of interest recently, but there is no clear picture regarding their functional importance. New findings by researchers at Karolinska Institutet suggest that despite the relative abundance and differential expression, a substantial number of circular RNAs may represent “passenger molecules” with little impact on a cell, reflecting a stochasticity of the gene expression and splicing processes.
In a new study at the Department of Biosciences and Nutrition (BioNut), Karolinska Institutet, the role of circular RNAs (circRNAs) have been addressed in the context of Hedgehog signaling-mediated medulloblastoma tumors. Medulloblastomas are the most common pediatric cancers in the brain, with the Sonic Hedgehog (SHH) molecular subtype as one of the largest, comprising about 30% of all medulloblastoma cases.
In the study, Ani Azatyan, Peter Zaphiropoulos and others have interrogated the expression profile of circular RNAs in Hedgehog signaling-dependent cell lines and medulloblastoma cells. This has been achieved by implementation of modified RNA-seq protocols to detect abundantly and/or differentially expressed circRNAs. Using stringent criteria, they identified seven circular RNAs with reduced expression in Hedgehog-dependent medulloblastoma versus cerebellum, with their corresponding mRNAs unchanged. However, the depletion of these circRNAs, as well as their overexpression did not have a significant impact on medulloblastoma cell proliferation based on two independent assays.
These findings highlight the complexity of gene expression outcomes and the possibility that gene products may not necessarily exert an obvious phenotypic impact in the cellular context where they are present. However, this does not exclude the possibility that some RNA circles may indeed act as "driver molecules" in specific settings.
Currently Azatyan and Zaphiropoulos are engaging into an extended analysis of a much larger cohort that includes samples from all four medulloblastoma subtypes, i.e., SHH, WNT, group-3 and group-4, which may allow to pinpoint roles for circRNAs during medulloblastoma development.
